Why data scientists shouldn’t need to know Kubernetes
[Hacker News discussion, Twitter thread]
Recently, there have been many heated discussions on what the job of a data scientist should entail [1, 2, 3]. Many companies expect data scientists to be full-stack, which includes knowing lower-level infrastructure tools such as Kubernetes (K8s) and resource management.
This post is to argue that while it’s good for data scientists to own the entire stack, they can do so without having to know K8s if they leverage a good infrastructure abstraction tool that allows them to focus on actual data science instead of getting YAML files to work.
The post starts with a hypothesis that the expectation for full-stack data scientists comes from the fact that their development and production environments are vastly different. It continues to discuss two steps of the solutions to bridge the gap between these two environments: the first step is containerization and the second step is infrastructure abstraction.
While containerization is more or less well-understood, infrastructure abstraction is a relatively new category of tools, and many people still confuse them with workflow orchestrations. The last part of the post is a comparison of various workflow orchestration and infrastructure tools, including Airflow, Argo, Prefect, Kubeflow, and Metaflow.
Roles and Responsibilities:
— Nick Heitzman 📊📈 (@NickDoesData) February 12, 2019
- Automate horrible business practices
- Write ad hoc SQL as needed
REQUIRED EXPERIENCE:
- 15 years exp deep learning in Python
- PhD thesis on Bayesian modeling
- NLP experience in 7 languages
- 10 years of creating Hadoop clusters from scratch


Notes
- Production is a spectrum. For some teams, production means generating nice plots from notebook results to show to the business team. For other teams, production means keeping your models up and running for millions of users per day. In the first scenario, the production environment is similar to the development environment. The production environment mentioned in this post is closer to the one in the second scenario.
- This post isn’t to argue whether K8s is useful. K8s is useful. In this post, we only discuss whether a data scientist needs to know K8s.
- This post isn’t to argue that it’s not useful to be full-stack. If you’re well-versed in every part of the pipeline, I know a dozen companies who would hire you on the spot (I’d try to recruit you if you let me). But you shouldn’t be expected to be full-stack to become a data scientist.
The full-stack expectations
About a year ago, I tweeted about a set of skills I thought was important to become an ML engineer or data scientist. The list covers almost every part of the workflow: querying data, modeling, distributed training, and setting up endpoints. It even includes tools like Kubernetes and Airflow.
Things I’d prioritize learning if I was to study to become a ML engineer again:
— Chip Huyen (@chipro) October 11, 2020
1. Version control
2. SQL + NoSQL
3. Python
4. Pandas/Dask
5. Data structures
6. Prob & stats
7. ML algos
8. Parallel computing
9. REST API
10. Kubernetes + Airflow
11. Unit/integration tests
The tweet seems to resonate with my audience. After that, Eugene Yan messaged me that he also wrote about how data scientists should be more end-to-end. Eric Colson, Stitch Fix’s Chief Algorithms Officer (who previously was also VP Data Science and Engineering at Netflix), wrote a post on “the power of the full-stack data science generalist and the perils of division of labor through function.”
When I wrote that tweet, I believed that Kubernetes was essential to the DS/ML workflow. This sentiment came from the frustration at my own job – my life as an ML engineer would’ve been much easier if I was more fluent with K8s.
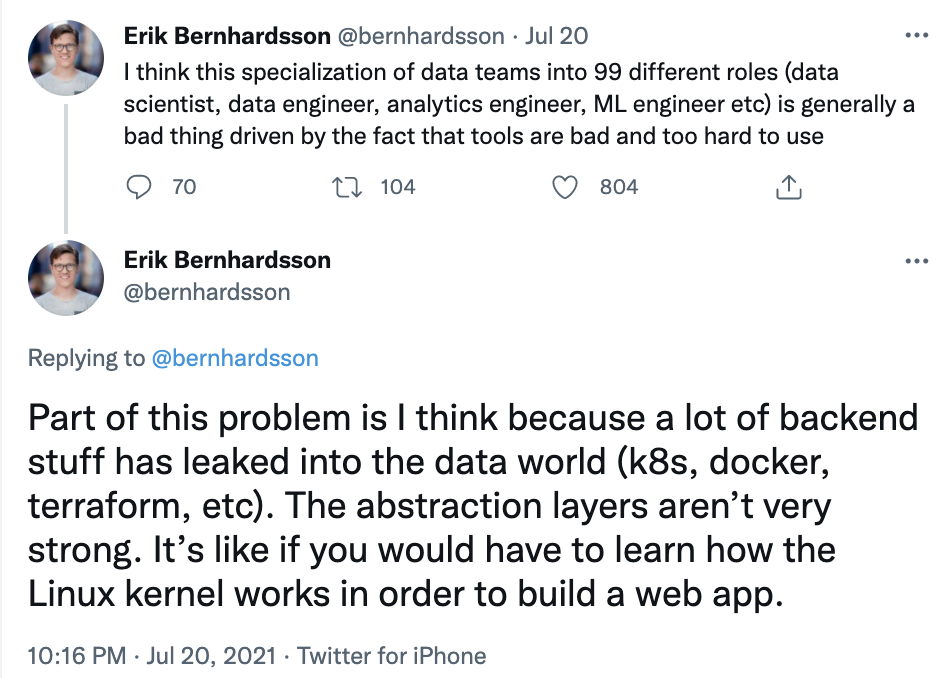
However, as I learned more about low-level infrastructure, I realized how unreasonable it is to expect data scientists to know about it. Infrastructure requires a very different set of skills from data science. In theory, you can learn both sets of skills. In practice, the more time you spend on one means the less time you spend on another. I love Erik Bernhardsson’s analogy that expecting data scientists to know about infrastructure is like expecting app developers to know about how Linux kernels work. I became a data scientist because I wanted to spend more time with data, not with spinning up AWS instances, writing Dockerfiles, scheduling/scaling clusters, or debugging YAML configuration files.
Development vs. production environments
So where does this unreasonable expectation come from?
One reason, in my opinion, is that there’s still a huge gap between the development (dev) and the production (prod) environments for data science. There are many differences between the dev and prod environments, but the two key differences that force data scientists to know two sets of tools for the two environments are scale and state.
| Development | Production | |
| Scale |
|
|
| State | Can keep on forever (a form of statefulness). Reproducible but inflexible.
|
Because instances are dynamically turned on/off, the setup is inherently stateless. Flexible but hard to reproduce.
|
During development, you might start a conda environment, work with notebooks, manipulate static data with pandas’ DataFrame, write model code with sklearn or PyTorch or TensorFlow, run and track multiple experiments.
Once you’re happy with the results (or you’ve run out of time), you pick the best model to productionize. Productionize your model basically means “move it from the development environment to the production environment”.
If you’re lucky, your production code can reuse the dev Python code and all you have to do is to copy and paste your notebook code into proper scripts. If you’re unlucky, you’ll have to rewrite your Python code into C++ or whatever language your company uses in production. Your dependencies (pandas, dask, PyTorch, TF, etc.) will need to be packaged and recreated on the instances your productionized models run on. If your model serves a lot of traffic and requires a lot of compute power, you might need to schedule your tasks. Previously, you’d have to manually spin up instances as well as shut them down when traffic is low, but most public cloud providers have taken care of this.
In traditional software development, CI/CD helps to close this gap. A well-developed set of tests can allow you to test how the changes you make locally will behave in production. However, CI/CD for data science is still falling short. On top of that, data distributions constantly shift in production. No matter how well your ML models do in development, you can’t be sure of how well they will do in production until they are actually in production.
Because of this separation, data science projects, therefore, involve two full sets of tools: one for the dev environment, and another for the prod environment.
[Bridging the gap] Part I: containerization
Container technology, including Docker, is designed to help you recreate dev environments in production. With Docker, you create a Dockerfile with step-by-step instructions on how to recreate an environment in which your model can run – e.g. install this package, download this pretrained model, set environment variables, navigate into a folder, etc. These instructions allow hardware anywhere to run your code.
If your application does anything interesting, you will likely need more than one container. Consider the case where your project consists of the featurizing code that is fast to run but requires a lot of memory, and the model training code that is slow to run but requires less memory. If you run both parts of the code on the same GPU instances, you’ll need GPU instances with high memory, which can be very expensive. Instead, you can run your featurizing code on CPU instances and the model training code on GPU instances. This means you’ll need a container for the featurizing instances and a container for the training instances.
Different containers might also be necessary when different steps in your pipeline have conflicting dependencies, such as your featurizer code requires NumPy 0.8 but your model requires NumPy 1.0.
When you have multiple containers with multiple instances, you might need to set up a network for them to communicate and share resources. You might also need a container orchestration tool to manage them and maintain high availability. Kubernetes is exactly that. It can help you spin up containers on more instances when you need more compute/memory as well as shutting down containers when you no longer need them.
Currently, to accommodate both the dev and prod environments, many teams choose one of the two following approaches:
-
Have a separate team to manage production
In this approach, the data science/ML team develops models in the dev environment. Then a separate team, usually the Ops/Platform/MLE team, to productionize the models in prod. This process has many drawbacks.
- Communication and coordination overhead: a team can become blockers for other teams. According to Frederick P. Brooks, what one programmer can do in one month, two programmers can do in two months.
- Debugging challenges: when something fails, you don’t know whether your team’s code or some other team’s code might have caused it. It might not have been because of your company’s code at all. You need cooperation from multiple teams to figure out what’s wrong.
- Finger-pointing: even when you’ve figured out what went wrong, each team might think it’s another team’s responsibility to fix it.
- Narrow context: no one has visibility into the entire process to optimize/improve the entire process. For example, the platform team has ideas on how to improve the infrastructure but they can only act on requests from data scientists, but data scientists don’t have to deal with infrastructure so they don’t care.
-
Data scientists own the entire process
In this approach, the data science team also has to worry about productionizing models. Data scientists become grumpy unicorns, expected to know everything about the process, and they might end up writing more boilerplate code than data science.
[Bridging the gap] Part II: infrastructure abstraction
What if we have an abstraction to allow data scientists to own the process end-to-end without having to worry about infrastructure?
What if I can just tell this tool: here’s where I store my data (S3), here are the steps to run my code (featurizing, modeling), here’s where my code should run (EC2 instances, serverless stuff like AWS Batch, Function, etc.), here’s what my code needs to run at each step (dependencies). And then this tool manages all the infrastructure stuff for me?
According to both Stitch Fix and Netflix, the success of a full-stack data scientist relies on the tools they have. They need tools that “abstract the data scientists from the complexities of containerization, distributed processing, automatic failover, and other advanced computer science concepts.”
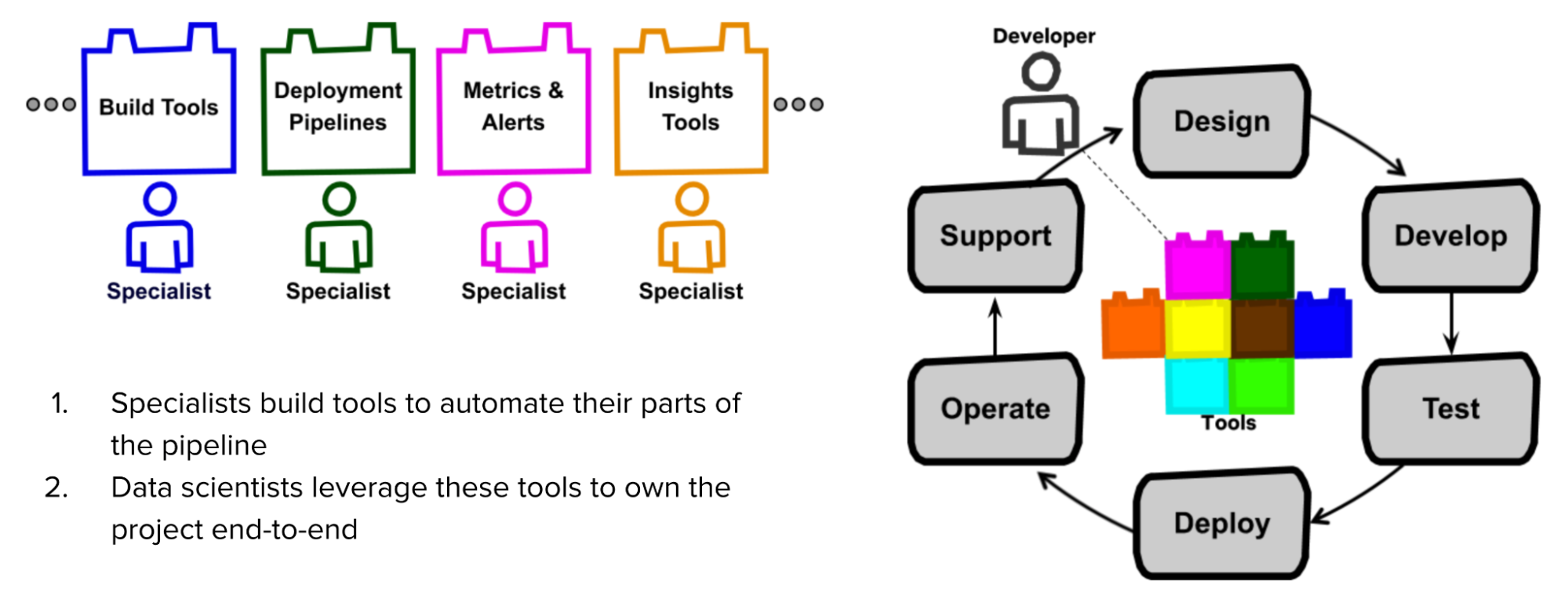
In Netflix’s model, the specialists – people who originally owned a part of the project – first create tools that automate their parts. Data scientists can leverage these tools to own their projects end-to-end.
The good news is that you don’t have to work for Netflix to have access to their tools. Two years ago, Netflix open-sourced Metaflow, an infrastructure abstraction tool that has allowed their data scientists to be full-stack without having to worry about low-level infrastructure.
The need for infrastructure abstraction for data science is a fairly recent problem for most companies. The main reason is that previously, most companies weren’t doing data science at a scale where infrastructure became a problem. Infrastructure abstraction is mainly useful when you have reasonably sophisticated cloud setups to abstract away from. The companies that benefit the most from it are the companies with a team of data scientists, non-trivial workflows, and multiple models in production.
Workflow orchestration vs. infrastructure abstraction
Because the need for infrastructure abstraction is a recent problem, the landscape for it is still open (and extremely confusing). If you’ve ever wondered what’s the difference between Airflow, Kubeflow, MLflow, Metaflow, Prefect, Argo, etc., you’re not alone. Paolo Di Tommaso’s awesome-pipeline repository features almost 200 workflow/pipeline toolkits. Most of them are workflow orchestration tools, not infrastructure abstraction, but because there is so much confusion around these two types of tools, let’s go over some of the key similarities and differences among them.
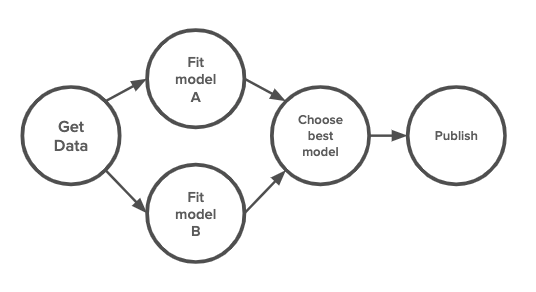
One reason for the confusion is that all these tools share the same fundamental concept. They all treat each workflow as a DAG, directed acyclic graph. Each step in the workflow is a node in the graph, and the edges between steps signal the order in which the steps should be executed. The differences lie in how the steps are defined, how they are packaged, and where they can be executed.
Workflow orchestration: Airflow vs. Prefect vs. Argo
Originally developed at Airbnb and released in 2014, Airflow is one of the earliest workflow orchestrators. It’s an amazing task scheduler that comes with a HUGE library of operators that makes it easy to use Airflow with different cloud providers, databases, storage options, etc. Airflow is a champion of the “configuration as code” principle. Its creators believed that data workflows are complex and should be defined using code (Python) instead of YAML or other declarative language. (And they’re right).
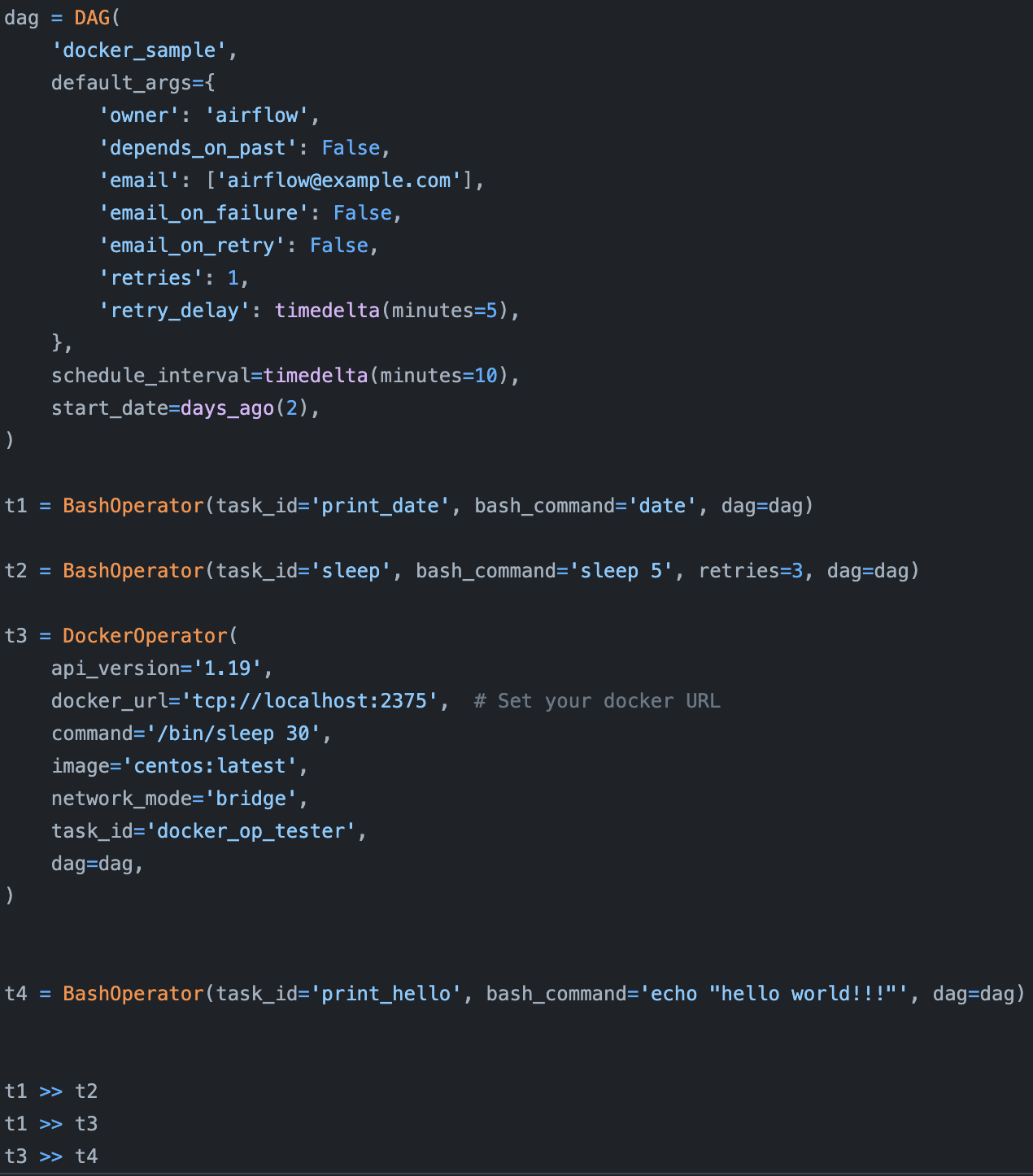
However, because Airflow was created earlier than most other tools, it had no tool to learn lessons from and suffers from many drawbacks, as discussed in detail in this blog post by Uber Engineering. Here, we’ll go over only three to give you an idea.
First, Airflow is monolithic, which means it packages the entire workflow into one container. If two different steps in your workflow have different requirements, you can, in theory, create different containers for them using Airflow’s DockerOperator, but it’s not that easy to do so.
Second, Airflow’s DAGs are not parameterized, which means you can’t pass parameters into your workflows. So if you want to run the same model with different learning rates, you’ll have to create different workflows.
Third, Airflow’s DAGs are static, which means it can’t automatically create new steps at runtime as needed. Imagine you’re reading from a database and you want to create a step to process each record in the database (e.g. to make a prediction), but you don’t know in advance how many records there are in the database, Airflow won’t be able to handle that.
The next generation of workflow orchestrators (Argo, Prefect) were created to address different drawbacks of Airflow.
Prefect’s CEO, Jeremiah Lowin, was a core contributor of Airflow. Their early marketing campaign drew intense comparison between Prefect and Airflow. Prefect’s workflows are parameterized and dynamic, a vast improvement compared to Airflow. It also follows the “configuration as code” principle so workflows are defined in Python.
However, like Airflow, containerized steps aren’t the first priority of Prefect. You can run each step in a container, but you’ll still have to deal with Dockerfiles and register your docker with your workflows in Prefect.
Argo addresses the container problem. Every step in an Argo’s workflow is run in its own container. However, Argo’s workflows are defined in YAML, which allows you to define each step and its requirements in the same file. But because of YAML, your workflow definitions can become messy and hard to debug.
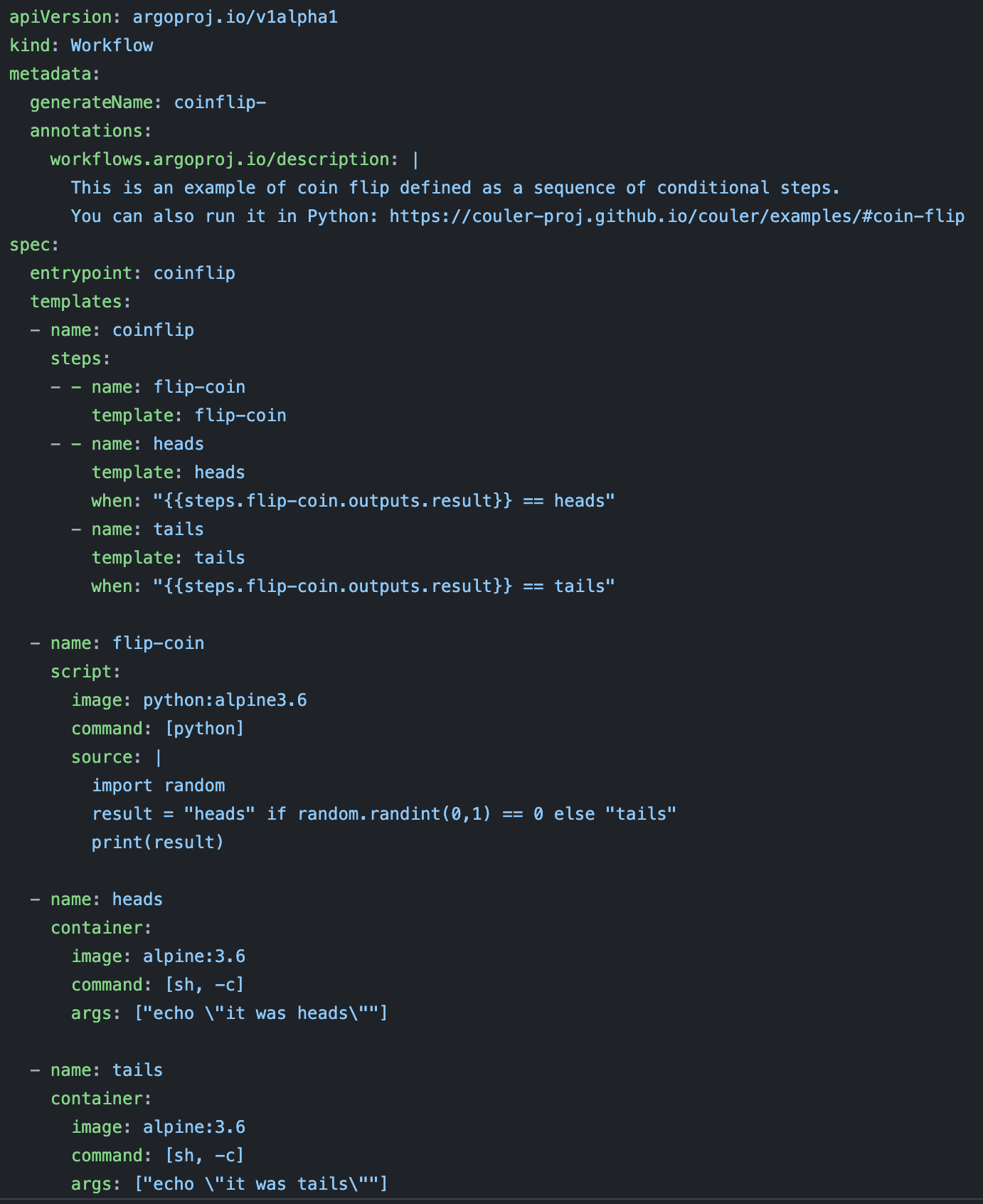
The main drawback of Argo, other than its messy YAML files, is that it can only run on Kubernetes clusters, which are only available in production. If you want to test the same workflow locally, you’ll have to use minikube or k3d.
Infrastructure abstraction: Kubeflow vs. Metaflow
Infrastructure abstraction tools like Kubeflow and Metaflow aim to help you run the workflow in both dev and prod environments by abstracting away infrastructure boilerplate code usually needed to run Airflow or Argo. They promise to give data scientists access to the full compute power of the prod environment from local notebooks, which effectively allows data scientists to use the same code in both dev and prod environments.
Even though they have some workflow orchestrating capacity, they are meant to be used with a bona fide workflow orchestrator. In fact, one component of Kubeflow is Kubeflow Pipelines, which is built on top of Argo.
Other than giving you a consistent environment for both dev and prod, Kubeflow and Metaflow provide some other nice properties.
- Version control: Automatically snapshot your workflow for model, data, and artifact.
- Dependency management: because they let you run each step of a workflow in its own container, you can manage dependencies for each step.
- Debuggability: When a step fails, you can resume your workflow from the failed step instead of from the start.
- Both are fully parameterized and dynamic.
Currently, Kubeflow is the more popular one because it integrates with K8s clusters (also, it was built by Google) while Metaflow only works with AWS services (Batch, Step Functions, etc.). However, it has recently spun out of Netflix to become a startup, so I expect it will grow to include a lot more use cases soon. At least, native K8s integration is in progress!
From a user experience perspective, IMO, Metaflow is superior. In Kubeflow, while you can define your workflow in Python, you still have to write a Dockerfile and a YAML file to specify the specs of each component (e.g. process data, train, deploy) before you can stitch them together in a Python workflow. So Kubeflow helps you abstract away other tools’ boilerplate by making you write Kubeflow boilerplate.
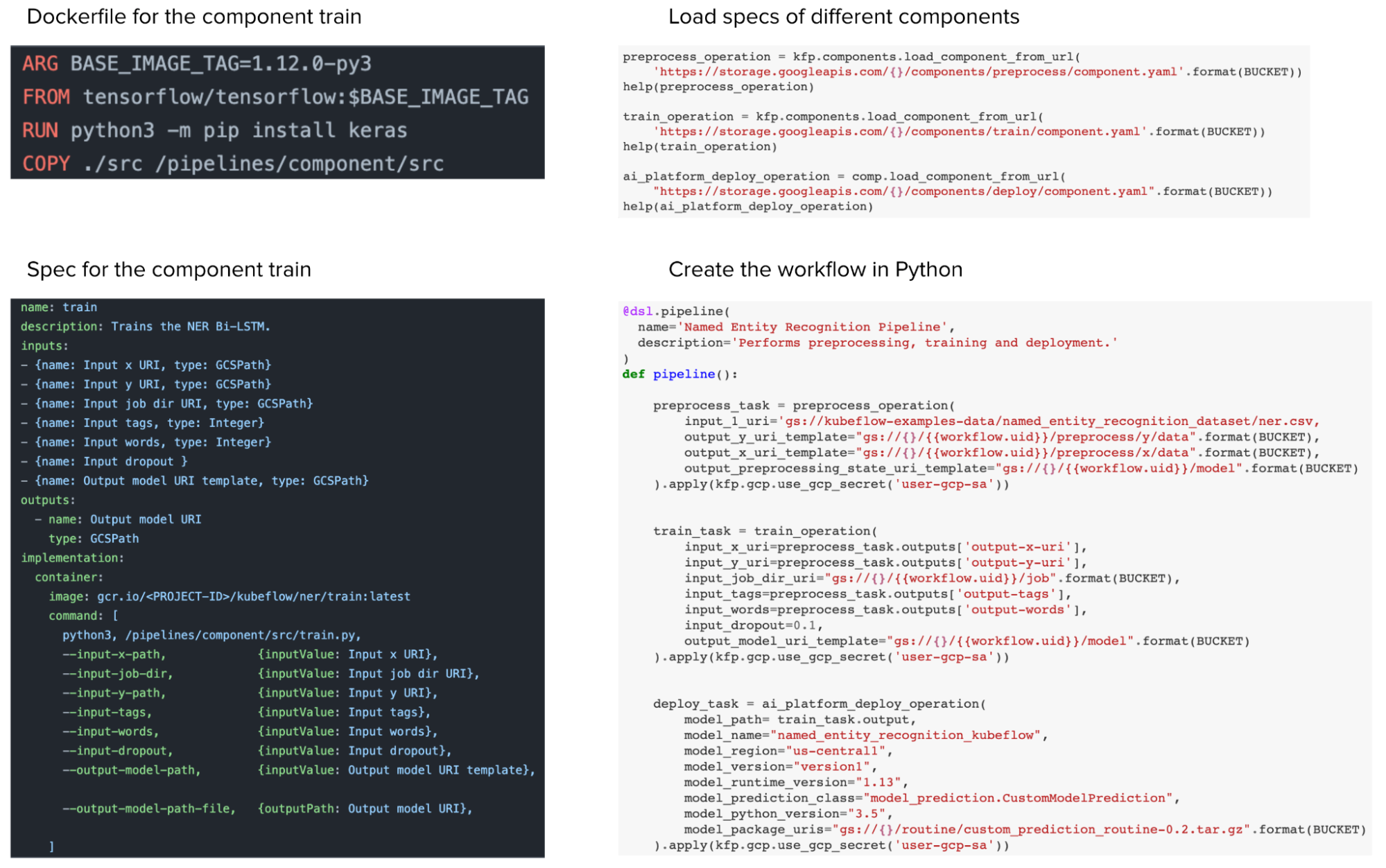
In Metaflow, you can use a Python decorator @conda to specify the requirements for each step — required libraries, memory & compute requirements – and Metaflow will automatically create a container with all these requirements to execute the step. You no longer have to write Dockerfiles or YAML files.
Metaflow allows you to work seamlessly with both dev and prod environments from the same notebook/script. You can run experiments with small datasets on local machines, and when you’re ready to run with the large dataset on the cloud, simply add @batch decorator to execute it on AWS Batch. You can even run different steps in the same workflow in different environments. For example, if a step requires a small memory footprint, it can run on your local machine. But if the next step requires a large memory footprint, you can just add @batch to execute it on the cloud.
# Example: sketch of a recommendation system that uses an ensemble of two models.
# Model A will be run on your local machine and model B will be run on AWS.
class RecSysFlow(FlowSpec):
@step
def start(self):
self.data = load_data()
self.next(self.fitA, self.fitB)
# fitA requires a different version of NumPy compared to fitB
@conda(libraries={"scikit-learn":"0.21.1", "numpy":"1.13.0"})
@step
def fitA(self):
self.model = fit(self.data, model="A")
self.next(self.ensemble)
@conda(libraries={"numpy":"0.9.8"})
# Requires 2 GPU of 16GB memory
@batch(gpu=2, memory=16000)
@step
def fitB(self):
self.model = fit(self.data, model="B")
self.next(self.ensemble)
@step
def ensemble(self, inputs):
self.outputs = (
(inputs.fitA.model.predict(self.data) +
inputs.fitB.model.predict(self.data)) / 2
for input in inputs
)
self.next(self.end)
def end(self):
print(self.outputs)
Takeaways
This post has become so much longer and denser than I intended. I blame it on the complexity and confusion of all the workflow-related tools, and on myself for not being able to find an easier way to explain them.
Here are some points that I hope you can come away with.
- The gap between the dev and production environments leads companies to expect data scientists to know two full sets of tools: one for the dev environment, one for the production environment.
- Owning a data science project end-to-end allows faster execution and lower communication overhead. However, it’d only make sense if we have good tools to abstract away lower level infrastructure to help data scientists focus on actual data science instead of configuration files.
- Infrastructure abstraction tools (Kubeflow, Metaflow) seem similar to workflow orchestrators (Airflow, Argo, Prefect) because they both treat workflows as DAGs. However, the main value of infrastructure abstraction is in allowing data scientists to use the code both locally and in production. You can use infrastructure abstraction tools together with workflow orchestrators.
- Infrastructure abstraction is one of those things that a lot of data scientists didn’t know they needed one until they used it. Try them out (Kubeflow is more involved but Metaflow takes 5 minutes to start).
Acknowledgments
Thanks Shreya Shankar for making this post a thousand times better. Thanks Hugo Bowne-Anderson, Luke Metz, Savin Goyal, Ville Tuulos, Ammar Asmro, Ramansh Sharma for being so patient with my never ending questions!
Updates
Yuan Tang, a top contributor to Argo, commented after the post:
- Argo is a large project, including Workflows, Events, CD, Rollouts, etc). It is more accurate to refer the sub-project Argo Workflows when comparing with other workflow engines.
- There are other projects that provide higher-level Python interface to Argo Workflows so that data scientists don’t have to work with YAML. Specifically, please check out Couler and Kubeflow Pipelines that use Argo Workflows as the workflow engine.
People have also mentioned other great tools that I haven’t been able to address here, such as MLFlow or Flyte. I’m still learning about the space. Your feedback is much appreciated. Thank you!